Pharmacogenomics of Heart Failure a Systematic Review Researchgate
Introduction
Multiple sclerosis (MS) is a chronic autoimmune disease characterized by the progressive infiltration of inflammatory cells to the central nervous system (CNS), demyelination and axonal damage. Although MS is affecting about 2.5 million individuals worldwide (1), its etiology remains largely unexplained. The clinical course of MS is highly heterogeneous, with current evidence suggesting that the combination of environmental and genetic run a risk factors is involved (1, ii). In full general, three types of MS accept been characterized, including a relapsing-remitting form of multiple sclerosis (RRMS) (eighty-85% of MS patients), which might evolve into secondary progressive MS (SPMS), and primary progressive MS (PPMS) manifesting in 15% of patients (3). Also, the response to the existing therapies largely varies betwixt individuals, with estimated non-responder rates ranging upwardly to l% for interferon beta (IFN-beta) and glatiramer acetate (GA) (4, v). Although the reasons for that variability remain unclear, several previous studies have implicated the role of genetics in response to MS handling (6, 7). While RRMS is the main focus of current pharmacogenomic research, equally information technology is the nearly common and the most responsive to electric current treatment options, only a few treatments are licensed to ho-hum the progressive form of the disease (three, 8). Approved medicines for MS include immunomodulatory and immunosuppressive drugs and monoclonal antibodies, including subcutaneous and intramuscular interferons, subcutaneous GA, intravenous (iv) natalizumab, oral fingolimod, teriflunomide and dimethyl fumarate, 4 mitoxantrone, iv alemtuzumab, and iv ocrelizumab, most of them clinically proven to be effective mainly in reducing annualized relapse rate (ARR) in the early stages of the disease (9). Among nigh widely prescribed get-go-line treatments worldwide remain IFN-beta and GA (10), which reduce frequency and severity of relapses in RRMS patients, subtract affliction progression rate and improve magnetic resonance imaging outcomes with minimal side furnishings. Those are the characteristics that are benign; notwithstanding, these drugs are just partially effective, and the response of individual patients to these therapies is highly unpredictable. Current literature suggests that approximately 30-fifty% of patients practice not respond well to kickoff-line therapies (depending on the response criteria used) (5), which is hypothesized to be in part attributed to inter-private genetic variability. In clinical exercise it is often the example that patients should fail to respond to beta-interferons or GA before receiving a second-line handling (9); moreover, clinical evaluation of response to the therapy requires 1-2 twelvemonth follow-up (11). It has previously been shown that there is a limited time window for effective intervention, during which the development of early brain atrophy, and thus cognitive and physical deficits, tin be minimized more effectively (12). Therefore, the biomarkers that would predict the responsiveness to therapy are indispensable to reduce adverse events and provide the maximized efficacy and condom early in the illness course.
Although pharmacogenomics in clinical do is increasingly available, at that place is currently no established genetic or whatsoever other clinical biomarker that would reliably predict a response of an private to selected MS therapy. However, with a growing number of approved treatment options for MS patients in contempo years and rapid advances in genomic technologies, personalized medicine has an opportunity to optimize handling for an individual.
In the nowadays commodity, we report the results of the conducted systematic review of currently published data on the pharmacogenomics of MS to review the electric current status of potential pharmacogenomic biomarkers and discuss their futurity potential in providing the most constructive treatment for an individual.
Methods
The systematic review was conducted according to Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) Statement guidelines1
Search Strategy
Manufactures on the pharmacogenomics of MS therapy published up to October ixth, 2018 were searched in the PubMed/MEDLINE database using the combinations of following keywords: multiple sclerosis, pharmacogenomics, pharmacogenetics, therapy response, genome-broad association report (GWAS), genome-wide, gene association study, candidate factor study, polymorphism/south, allele/south, and genetic variants. Search details are given in the Box 1. The search was express to articles published in the English linguistic communication. Firstly, articles were screened past title and abstract, next the full content was evaluated for their eligibility. The selection of manufactures and eligibility evaluation were carried out independently by the first two authors (KH and SR). We discussed discrepancies betwixt authors and reached an understanding on the selection of articles for systematic review. Finally, the main review articles were screened for possible boosted publications.
Box 1. Search details using PubMed database
(("multiple sclerosis"[All Fields] OR "interferon beta"[All Fields] OR "glatiramer acetate"[All Fields] OR "natalizumab"[All Fields] OR "fingolimod"[All Fields] OR "teriflunomide"[All Fields] OR "dimethyl fumarate"[All Fields] OR "mitoxantrone"[All Fields] OR "alemtuzumab"[All Fields] OR "ocrelizumab"[All Fields]) AND ("pharmacogenomics"[All Fields] OR "pharmacogenetics"[All Fields] OR "pharmacogenetic"[All Fields] OR "pharmacogenomic"[All Fields]))
OR
(("multiple sclerosis"[All Fields] OR "interferon beta"[All Fields] OR "glatiramer acetate"[All Fields] OR "natalizumab"[All Fields] OR "fingolimod"[All Fields] OR "teriflunomide"[All Fields] OR "dimethyl fumarate"[All Fields] OR "mitoxantrone"[All Fields] OR "alemtuzumab"[All Fields] OR "ocrelizumab"[All Fields]) AND ("GWAS"[All Fields] OR "genome-wide"[All Fields] OR "gene association study"[All Fields] OR "polymorphism"[All Fields] OR "polymorphisms"[All Fields] OR "allele"[All Fields] OR "gene variant"[All Fields] OR "alleles"[All Fields]) AND ("handling response"[All Fields] OR "therapy response"[All Fields] OR "response to therap"[All Fields] OR (response[All Fields] AND ("interferon-beta"[MeSH Terms] OR "interferon-beta"[All Fields] OR ("interferon"[All Fields] AND "beta"[All Fields]) OR "interferon beta"[All Fields])) OR (response[All Fields] AND ("glatiramer acetate"[MeSH Terms] OR ("glatiramer"[All Fields] AND "acetate"[All Fields]) OR "glatiramer acetate"[All Fields])) OR (response[All Fields] AND ("mitoxantrone"[MeSH Terms] OR "mitoxantrone"[All Fields])) OR (response[All Fields] AND ("teriflunomide"[Supplementary Concept] OR "teriflunomide"[All Fields])) OR (response[All Fields] AND ("fingolimod hydrochloride"[MeSH Terms] OR ("fingolimod"[All Fields] AND "hydrochloride"[All Fields]) OR "fingolimod hydrochloride"[All Fields] OR "fingolimod"[All Fields])) OR (response[All Fields] AND ("dimethyl fumarate"[MeSH Terms] OR ("dimethyl"[All Fields] AND "fumarate"[All Fields]) OR "dimethyl fumarate"[All Fields])) OR (response[All Fields] AND ("natalizumab"[MeSH Terms] OR "natalizumab"[All Fields])))).
Inclusion and Exclusion Criteria
Studies were included if they investigated response or nonresponse to handling, defined every bit relapse rate, by expanded inability status scale (EDSS) score or the definition was based on magnetic resonance imaging (MRI), in the clan to genetic variability. We included bachelor studies investigating the pharmacogenomics of all currently approved affliction-modifying treatment options for MS patients. We excluded manufactures that: (one) were not written in the English language language, (2) were commodity evaluations, example reports, reviews, written report protocols, (three) were using animal model, prison cell lines, in silico studies, (iv) investigated response by measuring NAbs/IFN-beta antibodies or studies evaluating therapeutic response past other biochemical tests, (5) were gene expression studies, and (half-dozen) investigated adverse drug reactions, such as liver and cardiac injury, acute leukemia and progressive multifocal leukoencephalopathy.
Data Collection
Ii authors (KH and SR) independently extracted the following data from manufactures: outset author'south last name, year of publication, PMID number, sample size, ethnic backgrounds of patients, method, genes, and polymorphisms tested, effect, significant associations with corresponding P-values and conviction intervals, response criteria, the duration of the follow-up period and medication investigated. Finally, The Pharmacogenomics Knowledgebase (PharmGKB) was reviewed for possible clinically actionable variants in MS treatments and to search for the level of show of the existing MS pharmacogenomic biomarkers. Genes with detected significant associations were annotated for Gene Ontology (GO) molecular functions and biological process using the online PANTHERTM tool version 13.1 (thirteen).
Results
In the master search, we identified a total of 297 manufactures in the PubMed database. Afterward reviewing titles and abstracts, 229 articles were excluded for the reasons presented in Figure 1. Boosted xx studies were excluded afterwards total-length review, because they investigated handling response by gene expression (n = 9), past measuring Nab/IFN-beta antibodies or by other biochemical tests (n = vi), investigated adverse drug reactions (n = 3), the sample size was modest (less than 20) (n = i), and investigated response to intravenous immunoglobulin (IVIG) (n = 1). In total, 48 publications investigating the association between genetic variation and treatment response met our inclusion criteria and were included in the systematic review: 40 (83 %) studies investigated treatment response to IFN-beta (5 GWAS and 35 candidate gene studies), 9 (19 %) studies investigated treatment response to GA (one GWAS and 8 candidate gene studies). Amidst them, iv studies investigated the response to both medications; IFN-beta and GA. In addition, nosotros identified two candidate gene studies investigating the response to mitoxantrone and one response to natalizumab. No studies on the pharmacogenomics of newest classes of agents, such as dimethyl fumarate, teriflunomide or fingolimod were identified. Eleven variants with the level of show iii and influence on treatment efficacy were found in the PharmGKB database. Results from candidate gene studies were mostly not replicated, and studies were performed in different populations. Furthermore, genes previously assessed in candidate cistron studies showed very little overlap with the pregnant GWAS associations. Nevertheless, few consequent significant findings (P < 0.05) were reported in the candidate gene studies.

Figure one. Menstruation diagram of identification and pick of studies.
IFN-beta
Interferon-beta i is one of the well-nigh commonly prescribed affliction-modifying therapies for patients with MS. Interferons are endogenous regulatory cytokines that bind to specific IFN alpha/beta receptors establish on the surface of the cells of the immune system and consequently alter the expression of many genes, depending on cell type - the inflammatory cytokine synthesis is inhibited (IL-12, IL-17, IL-23), while the production of anti-inflammatory cytokines (IL-4, IL-10) increases, which provokes differentiation toward a CD4+ T helper cell type phenotype -Th2 immune response (xiv). Additionally, interferon reduces the expression of matrix metalloproteases, affects the expression of jail cell adhesion molecules located on the endothelial surface and on the activated T-cell surface, which results in reduced T-jail cell activation and reduced lymphocyte migration across the blood-brain barrier (BBB). The potential antiviral action of IFN-beta has also been proposed (fifteen).
Candidate Gene Studies IFN-beta
We identified 35 studies investigating the association between genetic variability and response to IFN-beta, four of them likewise investigating the response to GA. The details of the included studies are presented in Supplementary Tabular array 1. The selection of candidate genes in these studies was mainly based on the proposed mechanisms of action of IFN-beta, and in contempo years, studies accept besides been designed to validate the significant results obtained from genome-wide studies. Some examples of candidate genes investigated were: HLA class 2 genes, MXA, genes coding for interferon receptors IFNAR1, IFNAR2 and other interferon-stimulated response elements (ISREs), interferon gamma IFNG, chemokine receptor CCR5, genes related to the type I IFN and TLR pathways, genes coding for GABA and glutamate receptors, genes encoding cytokines and their receptors, innate design recognition receptors, antigens CD46 and CD58, CTLA4, HAPLN1, ACE and APOE cistron.
There are a express number of studies conducted on the same polymorphisms. Furthermore, among those, the results were largely inconsistent. Sixteen (46%) of included IFN-beta candidate gene studies failed to identify any significant association comparing genetic variation between responders to not-responders. Non-meaning associations were repeatedly reported within the HLA locus of class I and/or II (six times) (4, 16–20), in IFNAR1 and INFAR2 genes (two times) (21, 22), in APOE gene (ii times) (23, 24), in IRF5 gene (25, 26), and NLRP3 cistron (27, 28). Other non-pregnant associations included MXA (29), HAPLN (xxx), IFNL3 (31), IRF8 (26), and GPC5 (26) genes. However, some reproducible pregnant associations between IFN-beta response and genetic variability have also been detected.
Despite the negative clan results between polymorphisms located in the promoter region of the MXA gene and IFN-beta response reported by Weinstock-Guttman et al. (29), the significant association was repeatedly demonstrated by ii independent studies, which together comprised three different SNPs in MXA gene, including rs464138 AA (P < 0.0001, OR = half dozen.23 [95% CI, 2.77–14.03]), rs2071430 G allele (P = 0.015, OR = three.four [95% CI, 1.ane-eleven.4]), and rs17000900 GG (P = 0.018, OR = two.4 [95% CI, 1.1-5.4]) (32, 33). One of those studies, which investigated 100 ISREs-containing genes in association to IFN-beta response heterogeneity, additionally identified significant associations betwixt IFNAR1 rs55884088 (GT)due north repeat (P = 0.036), LMP7 rs2071543 C allele (P = 0.002, OR = 6.iv [95% CI, 1.eight-24.ane]), and CTSS rs1136774 C allele (P = 0.02, OR = 0.iv [95% CI, 0.2-0.eight]) (32). Another SNP located in the tertiary intron of the IFNAR1 cistron was additionally associated with response to IFN-beta in the written report of Sriram et al. (21), suggesting a modest association of rs1012334 A allele with relapse-complimentary condition (P = 0.030, OR = 0.nine [95% CI, 0.2-one.2]). Furthermore, IFNAR1 rs1012335 Thou allele was associated with positive IFN-beta handling response (34) and was additionally, in allelic combinations, suggested equally a mark of pick for IFN-beta treatment over GA (6).
Another repeatedly studied variation is a 32-base of operations pair (bp) deletion of the CCR5 cistron (CCR5*d, rs333). A significant association between CCR5 deletion allele and IFN-beta treatment response in MS patients was confirmed by three independent analyses. In the study of Kulakova et al. (6), CCR5*d was more frequently constitute in Russian MS patients with optimal response to IFN-beta and GA non-responders, while CCR5*w/w was enriched in IFN-beta non-responders and GA responders. In the related study, allelic combinations of (CCR5*d + IFNAR1*G + IFNB1*T/T) or (CCR5*d + IFNAR1*G + IFNG*T) were proven to be benign for IFN-beta treatment efficacy (34). A significant association between CCR5*d and IFN-beta handling response in MS patients was also detected in the Egyptian population by Karam et al. (35) (P = 0.01, OR = three.2 [95%-CI, 1.one–viii.8]).
Sure genotypes of IRF5 gene polymorphisms (rs2004640 TT, P = 0.0006, and rs47281420 AA, P = 0.0023) were reported to exert a poor pharmacological response to IFN-beta, with more T2 lesions detected (36). In terms of item polymorphic loci, the finding IRF5 rs2004640 was replicated in an independent population within the same report (P = 0.037). The study of Vandenbroeck et al. (25) identified the trend toward a greater T allele frequency for the variant of IRF5 rs3807306 polymorphism in responders (P = 0.09), whereas no bear witness of an clan for IRF5 rs4728142 was detected. Evidence that an AA genotype of IRF8 rs17445836 polymorphism influences outcome-free survival in IFN-beta treated subjects was likewise found (P = 0.017, OR = 0.45 [95% CI, 0.2-0.ix]) (xix). Reverse, the study conducted in a Danish cohort of patients past Sellebjerg et al. (26), failed to identify whatsoever association between polymorphisms located in IRF5 (rs2004640, rs3807306, rs4728142) and in IRF8 (rs13333054 and rs17445836) genes.
The number of studies attempting to validate or farther investigate the results of GWAS studies is limited. Polymorphous loci in the GPC5 gene were reproducible with candidate-gene study of Cénit et al. (rs10492503 AA, P = 0.018, OR = 3.0 [95% CI, 1.iii-6.6]; rs1411751 GG, P = 0.012, OR = 3.seven [95% CI, 1.v-9.iv]), and GWAS by Byun et al. (rs10492503, rs9301789) (37), while the candidate gene study conducted by Sellebjerg et al. (26) yielded non-significant upshot. The aim of the written report conducted past Bustamante et al. (38) was to further investigate the results of ii GWAS studies that highlighted the importance of genes playing part in price-like receptor (TLR) pathways, type I interferon (IFN)-induced genes, and genes coding for GABA and glutamate receptors. An investigation of 384 polymorphisms located in those genes, detected only two significant polymorphisms (rs2277302 in PELI3 gene, P = 0.008, and rs832032 in GABRR3, P = 0.006 gene). Overall association of polymorphisms located in these pathways was therefore non confirmed (38).
As the testify of polygenic nature of IFN-beta handling response, allelic combinations (JAK2-IL10RB-GBP1-PIAS1 and JAK2-IL10-CASP3) were detected as significant, while no significant clan of tested individual polymorphisms was found (39). In some other written report, MS patients with non-GCC haplotypes (rs1800896, rs1800871, rs1800872) of the IL10 cistron experienced fewer new MRI T1-dissimilarity enhancing lesions than patients with the GCC haplotype (40).
Other positive associations included: intronic polymorphism rs2542109 of the USP18 gene, TGFB1 rs1800469, TRAILR-1 rs20576, CD46 rs2724385, GPC5 rs10492503 and rs1411751 polymorphisms, polymorphic microsatellite located in the first intron of the IFNG factor, and CD58 rs12044852 polymorphism.
Meaning associations (P < 0.05) betwixt treatment response and IFN-beta, identified in at least ane study, are presented in Table one.
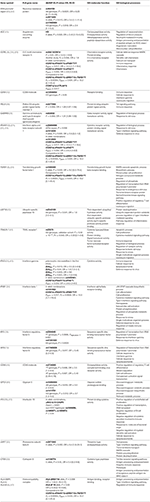
Table 1. Significant associations from candidate gene studies and IFN-beta MS treatment response along with selected gene ontology (GO) annotations. P perm , P-value permutation test; P f , P-value exact Fisher'southward exam.
GWAS Studies and IFN-beta
Currently, v GWAS studies investigating an association betwixt IFN-beta treatment response and genetic variation were carried out. GWAS study, which investigated SNPs in HLA- and not-HLA genes in association with the development of antibodies to IFN-beta therapy, was excluded from this review (48). None of the GWAS studies reported like results, but they uniquely suggested that multiple genes influence the handling response to IFN-beta. Furthermore, on the level of genes, well-nigh of the results were in deviation with previously conducted candidate-gene studies, thus providing novel candidate genes that might be involved in response to IFN-beta handling. However, it is important to notation that some potential candidate genes reported by contained GWAS studies were involved in the same biological pathways.
In the first GWAS study, conducted in 2008 by Byun et al. (37), authors found out that many of the detected differences between responders and non-responders were located in genes involved in ion channels and signal transduction pathways. Additionally, the authors too suggested that genetic variants in heparan sulfate proteoglycan genes (HAPLN1) might be useful as possible clinical predictors of response to MS therapy. Results of the second GWAS study conducted by Comabella et al. (11), indicated the importance of the glutamatergic system (GRIA3 factor) in patients response to IFN-beta therapy. The GWAS written report, conducted by Esposito and colleagues followed in 2015 and reported candidate intronic polymorphism rs9828519 in the SLC9A9 gene encoding for sodium/hydrogen exchanger establish in endosomes. For this gene, a broader role in MS pathogenesis, beyond treatment with IFN-beta, was also proposed. The gene product was functionally characterized to inhibit the development of pro-inflammatory CD4+ T cells (7).
In the study of Mahurkar et al. (49), none of the SNPs reached the level of genome-broad significance. The strongest associations were observed for FHIT factor and followed by variants in GAPVD1 and near ZNF697 gene. In the discovery stage of this report, samples were individually genotyped using Illumina® arrays, which distinguishes it from previously published GWAS studies where pooled genotyping was performed. A recent GWAS written report conducted past Clarelli et al. (fifty) investigated long-term handling response considering a iv-year follow-up report period and included just patients with extreme phenotypes of treatment responses. In contrast, all of the previous GWAS studies have taken into account 2-year follow up period. In summary, alterations in the genes involved in immunoregulatory processes, the glutamatergic system (GRIK2 and GRM3), and point transduction (GAPVD1) reached the highest significance.
Lack of overlap between GWAS studies likely reflects the differences in definitions of responders or not-responders, furthermore, GWAS studies covered populations of various ethnicities, including Italian, German, Spanish, and Australian. Also, the methodology was based on different genotyping platforms - kickoff two GWAS studies used Affymetrix genotyping platforms, covering 100 000 and 428 867 SNPs, respectively, while Illumina arrays were used in all of the afterwards studies (Illumina® Human 660-Quad platform, Illumina® 2.5M platform, Illumina® OmniExpress BeadChip and Illumina® OMNI-5M assortment). However, we showed that genes identified past GWAS studies were significantly enriched for ionotropic glutamate receptor signaling pathway (GO:0035235). Tiptop-ranking results from GWAS studies are summarized in Table 2.
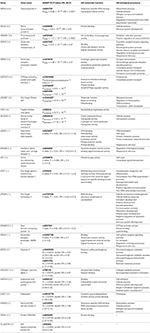
Table two. Genes with detected significant associations with response to IFN-beta in at least 1 written report and top-ranking results from GWAS studies along with selected factor ontology (Go) biological processes and molecular functions.
Candidate Factor Studies and Glatiramer Acetate
Glatiramer acetate is another widely prescribed disease-modifying therapy for patients with MS, with a circuitous and nonetheless not fully understood machinery of action. GA is a heterogeneous mixture of synthetic polymers made of random sequences of iv amino acids (51). Information technology acts through immunomodulatory actions to the cells of innate and caused immune response. Through binding to MHC Course II molecules, it participates in the generation of GA-specific T-cells and shifts their phenotype from pro-inflammatory helper-T types 1 and 17 (Th1/Th17) to anti-inflammatory regulatory T cells (Tregs) and helper-T type 2 (Th2) cells. Additionally, GA-specific T-cells are able to migrate through the BBB, where they induce local secretion of anti-inflammatory cytokines at the site of the lesions (51).
8 candidate gene studies investigating an association betwixt polymorphisms and treatment response to GA met our inclusion criteria. Four of them were mentioned above, every bit they as well investigated IFN-beta treatment response. Detailed data on studies is summarized in Supplementary Table ane. Hypothesis-driven approaches take primarily investigated genes involved in the mechanism of action of GA. Contrary to IFN-beta response, the HLA form I /II genes have been repeatedly positively associated with GA treatment response. The HLA DRB1 *1501 allele was demonstrated to influence the response to GA therapy in the cohort of 44 Italian RRMS patients (P = 0.008) (4) and in the cohort of 332 American patients, HLA-DBR1*1501/1501 genotype was significantly enriched among GA-responders (rs3135388, P AG/AA = 0.015, OR = 2.vii [95% CI, 1.2-6.0]) (19). The related report conducted in 64 American subjects with RRMS, showed that the presence of HLA DR15 or DQ6 alleles or the absenteeism of DR17 and DQ2 alleles is nominally associated with a favorable clinical response (52). The authors too demonstrated that the presence of the DR15-DQ6 haplotype and the absence of the DR17-DQ2 haplotype is significantly associated with positive treatment response (52). Furthermore, in a cohort of 296 Russian patients, the nominally significant clan of HLA-DRB1*4 allele with a positive response to GA was detected comparing responders to nonresponders and intermediate responders, P f = 0.015, OR = 2.02 [95% CI, 1.11–3.67] (53).
One of the beginning pharmacogenetics candidate-gene studies on GA, reported a significant association between GA response and ii SNPs, rs71878 in a T-cell receptor beta (TCRB) gene (P = 0.015, OR = 6.85 [95% CI, 1.45–31.9]) and rs2275235 in the cathepsin S (CTSS) (P = 0.014, OR = eleven.59 [95% CI, 1.6–81.9]) (54). Nominally pregnant associations were shown for boosted genes MBP, CD86, FAS, IL1R1, and IL12RB2. However, in the same experiment, no significant association for the HLA-DRB1*1501 allele was identified, suggesting the genetic heterogeneity of this region amidst the different populations equally the possible reason (54).
Using comparative pharmacogenomics approach investigating allelic combinations of CCR5, IFNAR1, TGFB1, DRB, and CTLA4 genes, the CCR5*westward/w genotype was the near enriched in GA responders compared to IFN-beta responders (half-dozen). In the most recent study examining association between GWAS identified MS susceptibility loci and efficacy of GA therapy in a Russian population of 296 RRMS patients, 5 SNPs were associated by themselves with result-gratis phenotype: EOMES rs2371108 T allele, CLEC16A rs6498169 A allele, IL22RA2 rs202573 GG genotype, PVT1 rs2114358 A allele, and HLA-DBR1*4 (P f = 0.032-0.00092). Authors demonstrated increased significance levels when taking into business relationship biallelic and triallelic combinations of these genes with additionally included polymorphic variants of TYK2, CD6, IL7RA and IRF8 genes (53).
Genes with at to the lowest degree one detected pregnant association (P < 0.05), along with Become biological processes and molecular functions, are presented in Tabular array 3.

Table 3. Genes with detected meaning associations with response to GA from candidate factor studies and GWAS studies along with selected factor ontology (GO) biological processes and molecular functions. P perm , P-value permutation test; P f , P-value exact Fisher'south exam.
GWAS and Glatiramer Acetate
To date, ane GWAS study investigating the research question of pharmacogenomics and GA has been published (56). Patients with extreme phenotypes were included in the assay, considering a 4-year follow-up period. Genotyping was conducted using Illumina OMNI-5M genome-wide array® covering 4301331 SNPs. Significant associations with treatment response were identified in the following genes: UVRAG (rs80191572), HLA-DQB2 (rs28724893), MBP (rs1789084), and ZAK (rs139890339). Marginal association with another polymorphism (rs470929) located in the MBP gene has previously been reported in the candidate gene report conducted by Grossman et al. (54). The MBP gene encodes the autoantigen myelin basic protein, which is attacked by the immune organisation in MS patients. Furthermore, GA was designed as an MBP mimetic (51). The results from the GWAS study warrant further confirmation in contained studies. Meaning results are presented in Table 3.
Mitoxantrone
Mitoxantrone is synthetic anthracenedione – a cytotoxic amanuensis that inhibits DNA repair via inhibition of topoisomerase II leading to a suppressed proliferation of T cells, B cells, and macrophages, decreased pro-inflammatory cytokine secretion, enhanced suppressor T jail cell role, and suppressed macrophage-mediated myelin degradation (57). Two studies investigating an association between genetic polymorphisms and mitoxantrone were published to date, providing conflicting results (58, 59). In the start study, authors proposed that SNPs in ABC-transporter genes (ABCB1 and ABCG2) might serve equally pharmacogenetic markers associated with clinical response to mitoxantrone therapy in patients with RRMS or SPMS forms of the disease. The second report failed to confirm the association in PPMS course of the affliction (59).
Natalizumab
Natalizumab is a humanized monoclonal antibody that inhibits the migration of lymphocytes via the BBB by inhibiting an adhesive molecule of anti-integrin-α4 (57). Currently, only one pharmacogenetic study in association to treatment response was conducted (60). Authors investigated an clan between polymorphisms in NQO1 and GSTP1 genes and handling efficacy. In a combined assay, it was constitute that patients who carried the wild-type genotype or only ane non-wild polymorphism for either gene have mayhap a better clinical outcome after receiving the natalizumab therapy.
PharmGKB Variants
Co-ordinate to PharmGKB levels of evidence for variant-drug associations, no clinically actionable variants with the level of evidence 1A or 1B exist for MS (October 9thursday, 2018). We identified eleven variant-drug combinations associated with treatment efficacy and with the level of show 3, which stands for an annotation based on a single significant (non nevertheless replicated) event or annotation for a variant-drug combination evaluated in multiple studies but lacking clear evidence of an association (61). The PharmGKB variants are presented in Table 4.

Table 4. Variant/gene-drug pairs currently listed in PharmGKB database (Oct 9th, 2018).
Neutralizing Antibodies (NAbs)
Part of the unresponsiveness to IFN-beta can too be explained by the development of neutralizing antibodies (NAbs) that reduce the drug efficacy. These develop in up to third of patients, depending on the IFN-beta product administered (62). Withal, the utilise of NAbs, as an early pharmacogenetic biomarker is limited considering NAbs develop only after half-dozen-24 months from initiation of treatment and patients may even revert to NAbs-negative over time (63). Additionally, NAbs-positivity explains the unresponsiveness to IFN-beta treatment just in a small proportion of patients (64). Nevertheless, it might be useful to include the information on NAbs in the pharmacogenetic studies of MS. Furthermore, the genetic markers that influence the development of NAbs take also been identified in patients with MS (candidate gene studies and GWAS) (48, 65–68).
Transcriptomic Pharmacogenetic Biomarkers
Although beyond the scope of this paper, several studies indicate that gene expression signatures could prove useful in predicting long-term treatment response in patients with MS (69–71). These studies revealed differences in the expression of genes related to IFN-beta signaling, TLR-4 signaling in monocytes, too every bit increased overall molecular response to IFN-beta in non-responders (72). Recently, RNA-sequencing in whole-blood showed that expression of a ribosomal poly peptide S6 was reduced in IFN-beta responders compared to non-responders (73). In another RNA-sequencing study, the different pre-treatment gene expression signature in peripheral blood mononuclear cells (PBMCs) was revealed in MS patients responsive to fingolimod compared to non-responders (74). However, near of the currently proposed transcriptomic biomarkers take merely moderate discriminative power and have yet not been validated (75, 76). Additionally, gene expression is more variable than genetic status and largely depends on various environmental factors, drugs co-administered (such as corticosteroids), specific cell populations studied (whole blood, PBMCs, T-cells), and differences in sampling times. Divergent findings can also be explained by the heterogeneity of technical protocols and clinical assessment of treatment response.
Word
In recent years, several actionable pharmacogenomics biomarkers have been identified, comprising many areas of medicine. The implementation of pharmacogenomics in clinical practice has, therefore, a smashing potential to enable more personalized handling with several benefits for patients and social club. Nevertheless, despite the increasing number of treatment options available to patients with MS and a high degree of variability in response to these treatments, there is still no reliable pharmacogenomic biomarker that would differentiate betwixt MS-treatment responders and non-responders. Since MS is a chronic progressive disorder, which requires life-long treatment, an early decision for the correct therapy may have a high clinical utility for MS patients. By choosing the right handling for a particular individual early in the disease grade, we can ho-hum downward the progression of the disease, avoid possible agin events and improve the efficiency of treatment.
Comprehensive systematic analysis of pharmacogenomic studies showed that the majority of the included studies (87.5%) are express to candidate genes, mostly hypothesized to exist involved in pathways of drug actions. We have observed that candidate gene studies largely lack the replication and confirmation of the results. However, nosotros take identified some genes, the variability of which has been investigated repeatedly, such as MXA, CCR5, GPC5, IFNAR1, IFNAR2, IRF5, NLRP3 genes, and HLA-region. The results of currently published candidate gene studies were mostly inconsistent, which may in part reflect the various report designs, including the inconsistent approach of defining response to treatment, too every bit limited sample sizes with bereft event size. Nevertheless, it is evident that biological processes defined past statistically meaning genes implicated in IFN-beta response are mostly immune-related and include regulation of interleukins production, positive regulation of regulatory T cell differentiation, negative regulation of cytokine product, type I interferon signaling pathway, mononuclear cell proliferation, cellular response to lipopolysaccharide, cellular response to interferon-gamma, regulation of cytokine-mediated signaling pathway, defense response to virus, leukocyte migration, and regulation of innate immune response.
Despite the proposed distinct immunomodulatory mechanisms of actions of IFN-beta and GA, we accept observed that some of the significant associations were identified in the same genes, or in genes involved in the same biological pathways. As an example, information technology has been found that the polymorphisms of the cathepsin Due south (CTSS) gene are associated with a response to the treatment of both IFN-beta and GA. Cathepsin S has cysteine-type peptidase activity and is involved in several biological processes, including Toll-like receptor signaling pathway, antigen processing and presentation of exogenous peptide antigen via MHC form Two, adaptive immune response, and proteolysis, also of homo myelin basic protein (MBP) (77). Furthermore, it has been suggested that discriminative allelic variants of the CCR5, IFNAR1, and TGFB1 genes, which are involved in MAPK cascade, defense force response, type I IFN signaling pathway, regulatory T jail cell differentiation, and apoptotic processes, may direct the handling decision between IFN-beta or GA (6).
In recent years, GWAS studies identified novel candidate genes, which remain to exist validated. Moreover, there was no overlap between the summit-ranked results of GWAS studies, which suggests that response to existing therapies is influenced by numerous polymorphisms in multiple genes. However, among potential candidate genes identified in GWAS studies of IFN-beta, we detected meaning enrichment for genes involved in the glutamate receptor-signaling pathway. Therefore, in the future, more global approaches, such equally GWAS or next-generation sequencing (NGS), are required to gain further insight into the pharmacogenomics of MS.
It is of import to admit the methodological heterogeneity betwixt the studies included in the present systematic review, such every bit variability of clinical characterization of the patients, differently defined clinical response, the varying duration of follow-up period amidst studies, and dissimilar genotyping platforms used in GWAS studies. It has previously been reported that the proportion of not-responders varies depending on the definition of treatment response used (78). The clinical criteria for phenotypic classification of patients (responders/non-responders) included: (1) relapse charge per unit (with different thresholds between studies), (ii) disease progression, which was about often measured past EDSS score, and (three) changes in MRI action, such as increase in T2 lesion brunt or T1 gadolinium (Gd) enhancing lesions on MRI. The detailed data on the definition of responders/non-responders for each written report are presented in Supplementary Table 1. Similarly, the follow-up period ranged from vi months (in one report) to 1 yr in ane study, two years in the bulk of the included studies and to four years in two recent studies.
More contained studies investigating the association betwixt already proposed polymorphisms in genes, such as GRHL3, NINJ2, TBXAS1, GRM3, GRIK2, and SLC9A9 and treatment response are warranted to plant reliable and accurate pharmacogenomics predictors. Futurity studies need to include a larger number of subjects of various ethnicities. It is as well crucial to utilize uniform and precise definitions of treatment response, standardized duration of the follow-upwardly menses, and comprehensive clinical characterization of patients.
Also, the GWAS studies are limited to common variants. Of note, rare variants contribute a major part of pharmacogenetic variability (79). In contempo years, many important advances in sequencing technologies have been achieved that volition in futurity enable a more comprehensive motion-picture show of pharmacogenetic variability in MS patients. Farther studies should also consider rare variants obtained by NGS technologies, such as exome or genome sequencing data. To the best of our knowledge, no study investigating the rare variation in exome or genome sequencing information of MS patients in association to handling response has been published to date.
Furthermore, nosotros suppose that the phenotype of the response to an immunomodulatory pleiotropic therapy, such as IFN-beta and GA, is a sum of numerous contributing genetic factors that were not sufficiently simultaneously and combinatorially assessed by current written report designs and methodologies. More studies investigating cumulative effects of polymorphisms in multiple genes (condiment effects or epistatic interactions), such as studies of Kulakova et al. (34, 53), are needed to gain a more than comprehensive insight into genetic variability in clan to the efficacy of treatment.
Another important aspect of future pharmacogenomics, peculiarly for the interpretation of rare variants, are publicly bachelor and easily updatable databases of pharmacogenomic variation, such as PharmGKB, CPIC, ClinVar, likewise as population-specific databases. Further standardized dosing recommendations and guidelines based on the patient's genomic test results are required, ideally integrated with demographic, phenotypic and clinical information.
Furthermore, most of the nerveless studies (94%) were conducted on patients treated with IFN-beta or GA. Lack of pharmacogenomic studies conducted on drugs approved in contempo years, such as dimethyl fumarate (Tecidifera®), teriflunomide (Aubagio®), and fingolimod (Gilenya®) limits the implementation of personalized medicine into clinical practise. An increasing number of new treatment options will in futurity enable more personalized treatment approaches; all the same, many genome-wide studies carried out on large sample sizes and in different populations are needed to reach reliable pharmacogenomics biomarkers for implementation into daily clinical practice.
In decision, electric current literature data suggests that genetic variability can significantly contribute to the response to treatment in patients with MS. In the hereafter, information technology is necessary to systematically evaluate the polymorphisms that were previously proposed to influence the response to treatment likewise every bit appraise the importance of rare variants and their effects on the treatment of MS. Boosted studies and larger ethnically homogenous cohorts are necessary to provide new insides and optimized use of MS drugs. More combinatorial written report designs are needed to assess the effect of several combinations of polymorphisms in diverse genes simultaneously to provide more than relevant information for the clinical employ of pharmacogenomics. Studies investigating the pharmacogenomics of newer medicines for MS are too necessary, using the articulate and uniform criteria of defining treatment response. Nosotros believe that all of the in a higher place, along with the rapid development of new medications and advances in genomic technologies, volition in future enable a more than personalized arroyo to MS treatment.
Writer Contributions
BP, SR, and KH contributed formulation and pattern of the study; KH and SR searched the database for potentially eligible articles and extracted the information. KH wrote the original typhoon. SR and BP contributed to manuscript revision. All authors reviewed the final version of the manuscript prior to submission for publication.
Funding
This study was funded by the Slovenian Research Agency (ARRS), grant no. P3-0326
Disharmonize of Interest Statement
The authors declare that the inquiry was conducted in the absence of whatsoever commercial or financial relationships that could exist construed every bit a potential conflict of interest.
Supplementary Material
The Supplementary Material for this article can be found online at: https://world wide web.frontiersin.org/manufactures/x.3389/fneur.2019.00134/total#supplementary-fabric
Abbreviations
ARR, annualized relapse rate; BBB, blood-encephalon barrier; CNS, primal nervous system; CPIC, Clinical Pharmacogenetics Implementation Consortium; EDSS, expanded disability status calibration; GA, glatiramer acetate; Become, gene ontology; GWAS, genome-wide association written report; IFN-beta, interferon beta; MHC, major histocompatibility circuitous; MRI, magnetic resonance imaging; MS, multiple sclerosis; Nabs, neutralizing antibodies; PBMCs, peripheral blood mononuclear cells; PharmGKB, The Pharmacogenomics Knowledgebase; PPMS, primary progressive multiple sclerosis; PRISMA, Preferred Reporting Items for Systematic Reviews and Meta-Analyses; RRMS, relapsing-remitting multiple sclerosis; SPMS, secondary progressive MS.
Footnotes
References
2. Belbasis Fifty, Bellou V, Evangelou Due east, Ioannidis JPA, Tzoulaki I. Environmental chance factors and multiple sclerosis: an umbrella review of systematic reviews and meta-analyses. Lancet Neurol. (2015) 14:263–73. doi: 10.1016/S1474-4422(14)70267-4
PubMed Abstract | CrossRef Full Text | Google Scholar
4. Fusco C, Andreone V, Coppola G, Luongo V, Guerini F, Step E, et al. HLA-DRB1*1501 and response to copolymer-ane therapy in relapsing-remitting multiple sclerosis. Neurology (2001) 57:1976–ix. doi: x.1212/WNL.57.eleven.1976
PubMed Abstract | CrossRef Full Text | Google Scholar
5. Río J, Nos C, Tintoré Thou, Téllez Northward, Galán I, Pelayo R, et al. Defining the response to interferon-β in relapsing-remitting multiple sclerosis patients. Ann Neurol. (2006) 59:344–52. doi: 10.1002/ana.20740
PubMed Abstract | CrossRef Full Text
6. Kulakova OG, Tsareva EY, Lvovs D, Favorov AV, Boyko AN, Favorova OO. Comparative pharmacogenetics of multiple sclerosis: IFN-beta versus glatiramer acetate. Pharmacogenomics (2014) 15:679–85. doi: 10.2217/pgs.14.26
PubMed Abstract | CrossRef Full Text | Google Scholar
7. Esposito F, Sorosina M, Ottoboni L, Lim ET, Replogle JM, Raj T, et al. A pharmacogenetic study implicates SLC9a9 in multiple sclerosis affliction activity. Ann Neurol. (2015) 78:115-27. doi: x.1002/ana.24429
PubMed Abstract | CrossRef Full Text | Google Scholar
8. Tur C, Moccia M, Barkhof F, Chataway J, Sastre-Garriga J, Thompson AJ, et al. Assessing treatment outcomes in multiple sclerosis trials and in the clinical setting. Nat Rev Neurol. (2018) 14:75–93. doi: 10.1038/nrneurol.2017.171
PubMed Abstract | CrossRef Full Text | Google Scholar
9. Ransohoff R, Hafler D, Lucchinetti C. Multiple sclerosis - a serenity revolution. Nat Rev Neurol. (2015) 11:134–42. doi: ten.1038/nrneurol.2015.xiv
CrossRef Full Text | Google Scholar
10. Tsareva Due east, Kulakova O, Boyko A, Favorova O. Pharmacogenetics of multiple sclerosis: personalized therapy with immunomodulatory drugs. Pharmacogenet Genomics (2016) 26:103–15. doi: 10.1097/FPC.0000000000000194
PubMed Abstract | CrossRef Full Text | Google Scholar
11. Comabella M, Craig DW, Morcillo-Suarez C, Rio J, Navarro A, Fernandez M, et al. Genome-wide scan of 500 000 single-nucleotide polymorphisms amid responders and nonresponders to interferon beta therapy in multiple sclerosis. Arch Neurol. (2009) 66:972–8. doi: 10.1001/archneurol.2009.150
PubMed Abstruse | CrossRef Full Text | Google Scholar
12. Duquette P, Giacomini PS, Bhan V, Hohol Grand, Schecter R. Balancing early assailment against risk of progression in multiple sclerosis. Can J Neurol Sci. (2015) 43:33–43. doi: x.1017/cjn.2015.302
PubMed Abstruse | CrossRef Full Text | Google Scholar
13. Mi H, Huang X, Muruganujan A, Tang H, Mills C, Kang D, et al. PANTHER version 11: expanded annotation data from gene ontology and reactome pathways, and data analysis tool enhancements. Nucleic Acids Res. (2017) 45:D183–nine. doi: 10.1093/nar/gkw1138
PubMed Abstract | CrossRef Full Text | Google Scholar
sixteen. Villoslada P, Barcellos LF, Rio J, Begovich AB, Tintore M, Sastre-Garriga J, et al. The HLA locus and multiple sclerosis in Spain. Role in illness susceptibility, clinical course and response to interferon-beta. J Neuroimmunol. (2002) 130:194–201. doi: 10.1016/S0165-5728(02)00215-1
PubMed Abstruse | CrossRef Total Text | Google Scholar
17. Fernández O, Fernández V, Mayorga C, Guerrero M, León A, Tamayo JA, et al. HLA class 2 and response to interferon-beta in multiple sclerosis. Acta Neurol Scand (2005) 112:391–4. doi: 10.1111/j.1600-0404.2005.00415.ten
PubMed Abstruse | CrossRef Full Text
18. Comabella M, Fernández-Arquero G, Río J, Guinea A, Fernández M, Cenit MC, et al. HLA class I and II alleles and response to handling with interferon-beta in relapsing-remitting multiple sclerosis. J Neuroimmunol. (2009) 210:116–nine. doi: 10.1016/j.jneuroim.2009.01.012
PubMed Abstruse | CrossRef Full Text | Google Scholar
xix. Gross R, Healy BC, Cepok Due south, Chitnis T, Khoury SJ, Hemmer B, et al. Population structure and HLA DRB1*1501 in the response of subjects with multiple sclerosis to starting time-line treatments. J Neuroimmunol. (2011) 233:168–74. doi: 10.1016/j.jneuroim.2010.ten.038
PubMed Abstract | CrossRef Full Text | Google Scholar
20. Samadzadeh S, Tabibian E, Sabokbar T, Shakoori A, Dehgolan SR, Armaki SA, et al. HLA-DRB1 does not accept a role in clinical response to interferon-beta among Iranian multiple sclerosis patients. J Neurol Sci. (2015) 352:37–forty. doi: 10.1016/j.jns.2015.03.004
CrossRef Full Text | Google Scholar
21. Sriram U, Barcellos LF, Villoslada P, Rio J, Baranzini SE, Caillier S, et al. Pharmacogenomic analysis of interferon receptor polymorphisms in multiple sclerosis. Genes Immun. (2003) 4:147–52. doi: x.1038/sj.gene.6363946
PubMed Abstract | CrossRef Full Text | Google Scholar
22. Leyva L, Fernández O, Fedetz One thousand, Blanco E, Fernández VE, Oliver B, et al. IFNAR1 and IFNAR2 polymorphisms confer susceptibility to multiple sclerosis simply not to interferon-beta handling response. J Neuroimmunol. (2005) 163:165–71. doi: x.1016/j.jneuroim.2005.02.010
CrossRef Full Text | Google Scholar
23. Carmona O, Masuet C, Alía P, Moral E, Alonso-Magdalena 50, Casado V, et al. Apolipoprotein alleles and the response to interferon-β-1b in multiple sclerosis. Eur Neurol. (2011) 65:132–7. doi: x.1159/000323982
PubMed Abstract | CrossRef Total Text | Google Scholar
24. Guerrero AL, Tejero MA, Gutiérrez F, Martín-Polo J, Iglesias F, Laherran East, et al. Influence of APOE cistron polymorphisms on interferon-beta treatment response in multiple sclerosis. Neurol (English language Ed (2011) 26:137–142. doi: x.1016/j.nrl.2010.06.003
PubMed Abstract | CrossRef Total Text | Google Scholar
25. Vandenbroeck Chiliad, Alloza I, Swaminathan B, Antigüedad A, Otaegui D, Olascoaga J, et al. Validation of IRF5 as multiple sclerosis adventure gene: putative role in interferon beta therapy and homo canker virus-vi infection. Genes Immun. (2011) 12:xl–45. doi: 10.1038/factor.2010.46
PubMed Abstruse | CrossRef Full Text | Google Scholar
26. Sellebjerg F, Søndergaard HB, Koch-Henriksen North, Sørensen PS, Oturai AB. Prediction of response to interferon therapy in multiple sclerosis. Acta Neurol Scand. (2014) 130:268–75. doi: ten.1111/i.12269
PubMed Abstruse | CrossRef Full Text | Google Scholar
27. Malhotra S, Sorosina K, Río J, Peroni South, Midaglia L, Villar LM, et al. NLRP3 polymorphisms and response to interferon-beta in multiple sclerosis patients. Mult Scler J. (2018) 24:1507–10. doi: ten.1177/1352458517739137
PubMed Abstract | CrossRef Full Text | Google Scholar
28. Malhotra S, Rio J, Urcelay E, Nurtdinov R, Bustamante MF, Fernandez O, et al. NLRP3 inflammasome is associated with the response to IFN-beta in patients with multiple sclerosis. Brain (2015) 138:644–52. doi: x.1093/brain/awu388
PubMed Abstract | CrossRef Total Text | Google Scholar
29. Weinstock-Guttman B, Tamaño-Blanco M, Bhasi K, Zivadinov R, Ramanathan M. Pharmacogenetics of MXA SNPs in interferon-β treated multiple sclerosis patients. J Neuroimmunol. (2007) 182:236–9. doi: 10.1016/j.jneuroim.2006.x.011
PubMed Abstract | CrossRef Total Text | Google Scholar
30. Cénit MDC, Blanco-Kelly F, de las Heras V, Bartolomé Thou, de la Concha EG, Urcelay E, et al. Glypican 5 is an interferon-beta response factor: a replication written report. Mult Scler (2009) 15:913–7. doi: x.1177/1352458509106509
PubMed Abstract | CrossRef Total Text
31. Malhotra Due south, Morcillo-Suárez C, Brassat D, Goertsches R, Lechner-Scott J, Urcelay E, et al. IL28B polymorphisms are not associated with the response to interferon-beta in multiple sclerosis. J Neuroimmunol. (2011) 239:101–4. doi: 10.1016/j.jneuroim.2011.08.004
CrossRef Full Text | Google Scholar
32. Cunningham S, Graham C, Hutchinson M, Droogan A, O'Rourke Thousand, Patterson C, et al. Pharmacogenomics of responsiveness to interferon IFN-β treatment in multiple sclerosis: a genetic screen of 100 type I interferon-inducible genes. Clin Pharmacol Ther. (2005) 78:635-46. doi: 10.1016/j.clpt.2005.08.018
PubMed Abstract | CrossRef Full Text | Google Scholar
33. Sayad A, Ghafouri-Fard S, Omrani MD, Noroozi R, Taheri M. Myxovirus resistance protein A (MxA) polymorphism is associated with IFNβ response in Iranian multiple sclerosis patients. Neurol Sci. (2017) 38:1093–9. doi: 10.1007/s10072-017-2935-four
PubMed Abstract | CrossRef Full Text | Google Scholar
34. Kulakova OG, Tsareva EY, Boyko AN, Shchur SG, Gusev EI, Lvovs D, et al. Allelic combinations of immune-response genes as possible composite markers of IFN-β efficacy in multiple sclerosis patients. Pharmacogenomics (2012) 13:1689–700. doi: 10.2217/pgs.12.161
PubMed Abstruse | CrossRef Full Text | Google Scholar
35. Karam RA, Rezk NA, Amer MM, Fathy HA. Immune response genes receptors expression and polymorphisms in relation to multiple sclerosis susceptibility and response to INF-β therapy. IUBMB Life (2016) 68:727–34. doi: ten.1002/iub.1530
PubMed Abstract | CrossRef Full Text | Google Scholar
36. Vosslamber Due south, van der Voort LF, van den Elskamp IJ, Heijmans R, Aubin C, Uitdehaag BMJ, et al. Interferon regulatory factor 5 gene variants and pharmacological and clinical result of Interferonβ therapy in multiple sclerosis. Genes Immun. (2011) 12:466–72. doi: 10.1038/gene.2011.eighteen
PubMed Abstract | CrossRef Full Text | Google Scholar
37. Byun E, Caillier SJ, Montalban X, Villoslada P, Fernández O, Brassat D, et al. Genome-broad pharmacogenomic analysis of the response to interferon beta therapy in multiple sclerosis. Arch Neurol. (2008) 65:337–44. doi: 10.1001/archneurol.2008.47
PubMed Abstract | CrossRef Full Text | Google Scholar
38. Bustamante MF, Morcillo-Suarez C, Malhotra S, Rio J, Leyva L, Fernandez O, et al. Pharmacogenomic report in patients with multiple sclerosis: responders and nonresponders to IFN-β. Neurol Neuroimmunol Neuroinflamm. (2015) 2:e154. doi: ten.1212/NXI.0000000000000154
PubMed Abstract | CrossRef Full Text | Google Scholar
39. O'Doherty C, Favorov A, Heggarty S, Graham C, Favorova O, Ochs M, et al. Genetic polymorphisms, their allele combinations and IFN-beta treatment response in Irish multiple sclerosis patients. Pharmacogenomics (2009) 10:1177–86. doi: 10.2217/pgs.09.41
PubMed Abstract | CrossRef Total Text | Google Scholar
xl. Wergeland South, Beiske A, Nyland H, Hovdal H, Jensen D, Larsen JP, et al. IL-x promoter haplotype influence on interferon handling response in multiple sclerosis. Eur J Neurol. (2005) 12:171–175. doi: 10.1111/j.1468-1331.2004.01102.x
PubMed Abstract | CrossRef Total Text | Google Scholar
41. Ristić S, Starčević Cizmarević N, Lavtar P, Lovrečić L, Perković O, Sepčić J, et al. Angiotensin-converting enzyme insertion/deletion gene polymorphism and interferon-β treatment response in multiple sclerosis patients. Pharmacogenet Genomics (2017) 27:232–v. doi: x.1097/FPC.0000000000000283
PubMed Abstruse | CrossRef Full Text
42. Torbati S, Karami F, Ghaffarpour M, Zamani 1000. Association of CD58 polymorphism with multiple sclerosis and response to interferon ß therapy in a subset of Iranian population. Cell J. 16:506–xiii. doi: 10.22074/cellj.2015.505
PubMed Abstract | CrossRef Total Text
43. Malhotra S, Morcillo-Suárez C, Nurtdinov R, Rio J, Sarro Eastward, Moreno M, et al. Roles of the ubiquitin peptidase USP18 in multiple sclerosis and the response to interferon-β treatment. Eur J Neurol. (2013) 20:1390–7. doi: x.1111/ene.12193
PubMed Abstract | CrossRef Full Text
44. López-Gómez C, Pino-Ángeles A, Órpez-Zafra T, Pinto-Medel MJ, Oliver-Martos B, Ortega-Pinazo J, et al. Candidate factor study of TRAIL and TRAIL receptors: association with response to interferon beta therapy in multiple sclerosis patients. PLoS O NE(2013) 8:e62540. doi: 10.1371/journal.pone.0062540
PubMed Abstruse | CrossRef Total Text
45. Martínez A, De Las Heras V, Mas Fontao A, Bartolomé Grand, De La Concha EG, Urcelay E, et al. An IFNG polymorphism is associated with interferon-beta response in Castilian MS patients. J Neuroimmunol. (2006) 173:196–nine. doi: 10.1016/j.jneuroim.2005.12.002
PubMed Abstract | CrossRef Full Text
46. Alvarez-Lafuente R, Blanco-Kelly F, Garcia-Montojo M, Martínez A, De Las Heras V, Dominguez-Mozo MI, et al. CD46 in a Spanish cohort of multiple sclerosis patients: genetics, mRNA expression and response to interferon-beta treatment. Mult Scler. (2011) 17:513–20. doi: 10.1177/1352458510393263
PubMed Abstract | CrossRef Total Text | Google Scholar
47. Mazdeh 1000, Taheri Yard, Sayad A, Bahram Due south, Omrani Dr., Movafagh A, et al. HLA genes as modifiers of response to IFN-beta-1a therapy in relapsing-remitting multiple sclerosis. Pharmacogenomics (2016) 17:489–98. doi: x.2217/pgs.xvi.2
PubMed Abstract | CrossRef Full Text | Google Scholar
48. Weber F, Cepok S, Wolf C, Berthele A, Uhr One thousand, Bettecken T, et al. Single-nucleotide polymorphisms in HLA- and non-HLA genes associated with the development of antibodies to interferon-β therapy in multiple sclerosis patients. Pharmacogenomics J. (2012) 12:238–45. doi: 10.1038/tpj.2011.xiv
PubMed Abstruse | CrossRef Full Text | Google Scholar
49. Mahurkar S, Moldovan Grand, Suppiah Five, Sorosina M, Clarelli F, Liberatore G, et al. Response to interferon-beta treatment in multiple sclerosis patients: a genome-wide association study. Pharmacogenomics J. (2017) 17:312–8. doi: 10.1038/tpj.2016.20
PubMed Abstruse | CrossRef Total Text | Google Scholar
50. Clarelli F, Liberatore G, Sorosina G, Osiceanu AM, Esposito F, Mascia E, et al. Pharmacogenetic study of long-term response to interferon-beta treatment in multiple sclerosis. Pharmacogenomics J. (2017) 17:84–91. doi: 10.1038/tpj.2015.85
PubMed Abstract | CrossRef Full Text | Google Scholar
51. Hasson T, Kolitz S, Towfic F, Laifenfeld D, Bakshi S, Beriozkin O, et al. Functional effects of the antigen glatiramer acetate are circuitous and tightly associated with its composition. J Neuroimmunol. (2016) 290:84–95. doi: x.1016/j.jneuroim.2015.11.020
PubMed Abstract | CrossRef Total Text | Google Scholar
52. Dhib-Jalbut S, Valenzuela RM, Ito G, Kaufman M, Ann Picone M, Buyske S. HLA DR and DQ alleles and haplotypes associated with clinical response to glatiramer acetate in multiple sclerosis. Mult Scler Relat Disord. (2013) ii:340–eight. doi: 10.1016/j.msard.2013.02.005
PubMed Abstruse | CrossRef Total Text | Google Scholar
53. Kulakova O, Bashinskaya V, Kiselev I, Baulina Northward, Tsareva E, Nikolaev R, et al. Pharmacogenetics of glatiramer acetate therapy for multiple sclerosis: the impact of genome-wide association studies identified disease risk loci. Pharmacogenomics (2017) 18:1563–74. doi: x.2217/pgs-2017-0058
PubMed Abstruse | CrossRef Full Text | Google Scholar
54. Grossman I, Avidan N, Singer C, Goldstaub D, Hayardeny L, Eyal E, et al. Pharmacogenetics of glatiramer acetate therapy for multiple sclerosis reveals drug-response markers. Pharmacogenet Genomics (2007) 17:657–66. doi: ten.1097/FPC.0b013e3281299169
PubMed Abstract | CrossRef Total Text | Google Scholar
55. Tsareva EY, Kulakova OG, Boyko AN, Shchur SG, Lvovs D, Favorov AV, et al. Allelic combinations of immune-response genes associated with glatiramer acetate handling response in Russian multiple sclerosis patients. Pharmacogenomics (2012) 13:43–53. doi: ten.2217/pgs.eleven.136
PubMed Abstract | CrossRef Full Text | Google Scholar
56. Ross CJ, Towfic F, Shankar J, Laifenfeld D, Thoma M, Davis G, et al. A pharmacogenetic signature of loftier response to Copaxone in belatedly-stage clinical-trial cohorts of multiple sclerosis. Genome Med. (2017) nine:one–15. doi: ten.1186/s13073-017-0436-y
PubMed Abstract | CrossRef Full Text | Google Scholar
57. Auricchio F, Scavone C, Cimmaruta D, Di Mauro G, Capuano A, Sportiello Fifty, et al. Drugs approved for the handling of multiple sclerosis: review of their safety profile. Skilful Opin Drug Saf. (2017) 16:1359–71. doi: 10.1080/14740338.2017.1388371
PubMed Abstruse | CrossRef Total Text | Google Scholar
58. Cotte S, Von Ahsen N, Kruse N, Huber B, Winkelmann A, Zettl U.k., et al. ABC-transporter cistron-polymorphisms are potential pharmacogenetic markers for mitoxantrone response in multiple sclerosis. Encephalon (2009) 132:2517–xxx. doi: 10.1093/brain/awp164
PubMed Abstract | CrossRef Total Text | Google Scholar
59. Grayness Née Cotte S, Salmen Née Stroet A, von Ahsen N, Starck M, Winkelmann A, Zettl UK, et al. Lack of efficacy of mitoxantrone in primary progressive Multiple Sclerosis irrespective of pharmacogenetic factors: a multi-middle, retrospective analysis. J Neuroimmunol. (2015) 278:277–9. doi: 10.1016/j.jneuroim.2014.11.017
CrossRef Full Text | Google Scholar
60. Alexoudi A, Zachaki South, Stavropoulou C, Gavrili S, Spiliopoulou C, Papadodima S, et al. Possible implication of GSTP1 and NQO1 polymorphisms on natalizumab response in multiple sclerosis. Ann Clin Lab Sci. (2016) 46:586–91.
PubMed Abstract | Google Scholar
61. Whirl-Carrillo M, McDonogh Eastward, Herbet J, Gong L, Sangkuhl K, Thotn C, et al. Pharmacogenomics knowledge for personlized medicine. Clin Pharmacol Ther. (2012) 92:414–vii. doi: x.1038/clpt.2012.96
PubMed Abstract | CrossRef Full Text
62. Bertolotto A, Deisenhammer F, Gallo P, Sölberg Sørensen Per. Immunogenicity of interferon beta: differences among products. J Neurol. (2004) 251 (Suppl. 2):II15–II24. doi: 10.1007/s00415-004-1204-7
PubMed Abstract | CrossRef Total Text | Google Scholar
63. Hartung HP, Freedman MS, Polman CH, Edan 1000, Kappos L, Miller DH, et al. Interferon β-1b-neutralizing antibodies v years after clinically isolated syndrome. Neurology (2011) 77:835–43. doi: ten.1212/WNL.0b013e31822c90d7
PubMed Abstruse | CrossRef Full Text | Google Scholar
64. Sbardella E, Tomassini 5, Gasperini C, Bellomi F, Cefaro LA, Morra VB, et al. Neutralizing antibodies explain the poor clinical response to Interferon beta in a small proportion of patients with Multiple Sclerosis: a retrospective study. BMC Neurol. (2009) 9:54. doi: 10.1186/1471-2377-9-54
PubMed Abstract | CrossRef Full Text | Google Scholar
65. Hoffmann S, Cepok Southward, Grummel V, Lehmann-Horn K, Hackermueller J, Stadler PF, et al. HLA-DRB1*0401 and HLA-DRB1*0408 are strongly associated with the evolution of antibodies against interferon-β therapy in multiple sclerosis. Am J Hum Genet. (2008) 83:219–27. doi: 10.1016/j.ajhg.2008.07.006
PubMed Abstruse | CrossRef Total Text | Google Scholar
66. Buck D, Cepok Southward, Hoffmann Due south, Grummel V, Jochim A, Berthele A, et al. Influence of the HLA-DRB1 genotype on antibody development to interferon beta in multiple sclerosis. Arch Neurol. (2011) 68:480–7. doi: x.1001/archneurol.2011.65
PubMed Abstract | CrossRef Full Text | Google Scholar
67. Núñez C, Cénit MC, Alvarez-Lafuente R, Río J, Fernández-Arquero K, Arroyo R, et al. HLA alleles as biomarkers of high-titre neutralising antibodies to interferon-β therapy in multiple sclerosis. J Med Genet. (2014) 51:395–400. doi: 10.1136/jmedgenet-2014-102348
PubMed Abstract | CrossRef Full Text | Google Scholar
68. Link J, Ryner ML, Fink Grand, Hermanrud C, Lima I, Brynedal B, et al. Human leukocyte antigen genes and interferon beta preparations influence run a risk of developing neutralizing anti-drug antibodies in multiple sclerosis. PLoS ONE (2014) ix:e90479. doi: ten.1371/periodical.pone.0090479
PubMed Abstract | CrossRef Full Text | Google Scholar
69. van Baarsen LGM, Vosslamber S, Tijssen M, Baggen JMC, van der Voort LF, Killestein J, et al. Pharmacogenomics of interferon-β therapy in multiple sclerosis: baseline IFN signature determines phamacological differences betwixt patients. PLoS Ane (2008) three:e1927. doi: 10.1371/journal.pone.0001927
PubMed Abstract | CrossRef Full Text | Google Scholar
70. Comabella Thousand, Lünemann JD, Río J, Sánchez A, López C, Julià E, et al. A type i interferon signature in monocytes is associated with poor response to interferon-β in multiple sclerosis. Brain (2009) 132:3353–65. doi: x.1093/brain/awp228
PubMed Abstract | CrossRef Full Text | Google Scholar
71. Goertsches RH, Zettl UK, Hecker One thousand. Sieving treatment biomarkers from claret gene-expression profiles: A pharmacogenomic update on ii types of multiple sclerosis therapy. Pharmacogenomics (2011) 12:423–32. doi: ten.2217/pgs.10.190
PubMed Abstract | CrossRef Full Text | Google Scholar
72. Rudick RA, Rani MRS, Xu Y, Lee JC, Na J, Shrock J, et al. Excessive biologic response to IFNβ is associated with poor treatment response in patients with multiple sclerosis. PLoS ONE (2011) half dozen:e19262. doi: 10.1371/journal.pone.0019262
PubMed Abstract | CrossRef Full Text | Google Scholar
73. Parnell GP, Gatt PN, McKay FC, Schibeci S, Krupa M, Powell JE, et al. Ribosomal protein S6 mRNA is a biomarker upregulated in multiple sclerosis, downregulated by interferon treatment, and affected past season. Mult Scler J. (2014) 20:675–85. doi: 10.1177/1352458513507819
PubMed Abstract | CrossRef Full Text | Google Scholar
74. Moreno-Torres I, González-García C, Marconi One thousand, García-Grande A, Rodríguez-Esparragoza L, Elvira Five, et al. Immunophenotype and transcriptome profile of patients with multiple sclerosis treated with fingolimod: Setting up a model for prediction of response in a two-year translational study. Forepart Immunol. (2018) nine:1693. doi: 10.3389/fimmu.2018.01693
PubMed Abstract | CrossRef Full Text | Google Scholar
75. Hecker M, Paap BK, Goertsches RH, Kandulski O, Fatum C, Koczan D, et al. Reassessment of claret gene expression markers for the prognosis of relapsing-remitting multiple sclerosis. PLoS ONE (2011) vi:e29648. doi: ten.1371/journal.pone.0029648
PubMed Abstract | CrossRef Full Text | Google Scholar
76. Martire Southward, Navone ND, Montarolo F, Perga S, Bertolotto A. A gene expression written report denies the ability of 25 candidate biomarkers to predict the interferon-beta treatment response in multiple sclerosis patients. J Neuroimmunol. (2016) 292:34–ix. doi: 10.1016/j.jneuroim.2016.01.010
PubMed Abstract | CrossRef Full Text | Google Scholar
77. Brook H, Schwarz G, Schröter CJ, Deeg Thousand, Baier D, Stevanovic S, et al. Cathepsin S and an asparagine-specific endoprotease dominate the proteolytic processing of human being myelin basic protein in vitro. Eur J Immunol. (2001) 31:3726–36. doi: 10.1002/1521-4141(200112)31:12<3726::AID-IMMU3726>iii.0.CO;2-O
PubMed Abstract | CrossRef Full Text | Google Scholar
78. Río J, Comabella M, Montalban X. Predicting responders to therapies for multiple sclerosis. Nat Rev Neurol. (2009) 5:553–lx. doi: 10.1038/nrneurol.2009.139
PubMed Abstract | CrossRef Full Text
79. Nelson MR, Wegmann D, Ehm MG, Kessner D, St P, Verzilli C, et al. NIH Public Access. (2015) 337:100–104. doi: ten.1126/scientific discipline.1217876
CrossRef Full Text
boltonyousterromme.blogspot.com
Source: https://www.frontiersin.org/articles/10.3389/fneur.2019.00134/full
0 Response to "Pharmacogenomics of Heart Failure a Systematic Review Researchgate"
Publicar un comentario